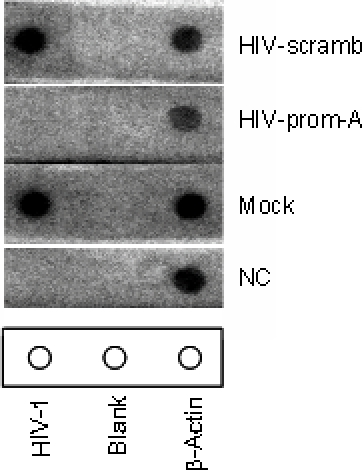
Nuclear run-off assay. (A) Diagram of the nuclear run-off assay. (B)... | Download Scientific Diagram
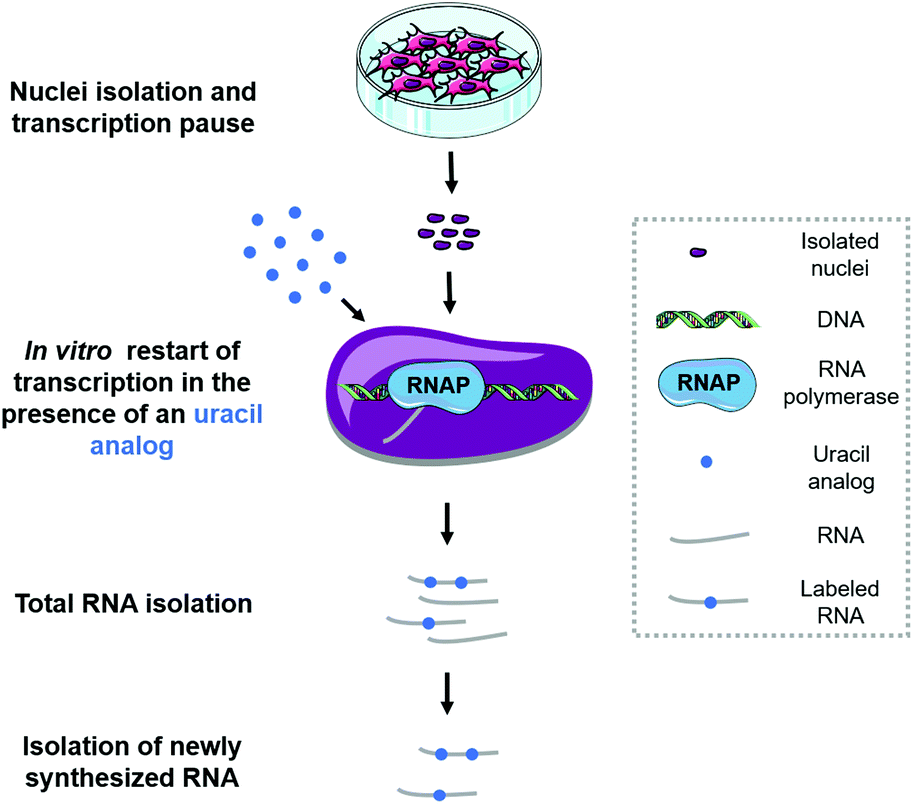
Methods for the analysis of transcriptome dynamics - Toxicology Research (RSC Publishing) DOI:10.1039/C9TX00088G

Analysis of Viral Promoter Usage in EBV-Infected Cell Lines: A Comparison of qPCR Following Conventional RNA Isolation and Nuclear Run-On Assay | SpringerLink

![PDF] Quantitative nuclear run-off transcription assay. | Semantic Scholar PDF] Quantitative nuclear run-off transcription assay. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/e96bfc026b1026d7b06d2cb4532a04c55ad9173d/1-Figure1-1.png)